11.11. Compute Feature Reference Misorientations
Group (Subgroup)
Statistics (Crystallography)
Description
This Filter calculates the misorientation angle between each Cell within a Feature and a reference orientation for that Feature. The user can choose the reference orientation to be used for the Features from a drop-down menu. The options for the reference orientation are the average orientation of the Feature or the orientation of the Cell that is furthest from the boundary of the Feature.
Note: the average orientation of the Feature is a typical choice, but if the Feature has undergone plastic deformation and the amount of lattice rotation developed is of interest, then it may be more reasonable to use the orientation near the center of the Feature as it may not have rotated and thus serve as a better reference orientation.
IPF Colors <001> Direction
This is the data set’s IPF colors by the <001> direction which shows the reader the relative orientation gradients within each feature (grain).

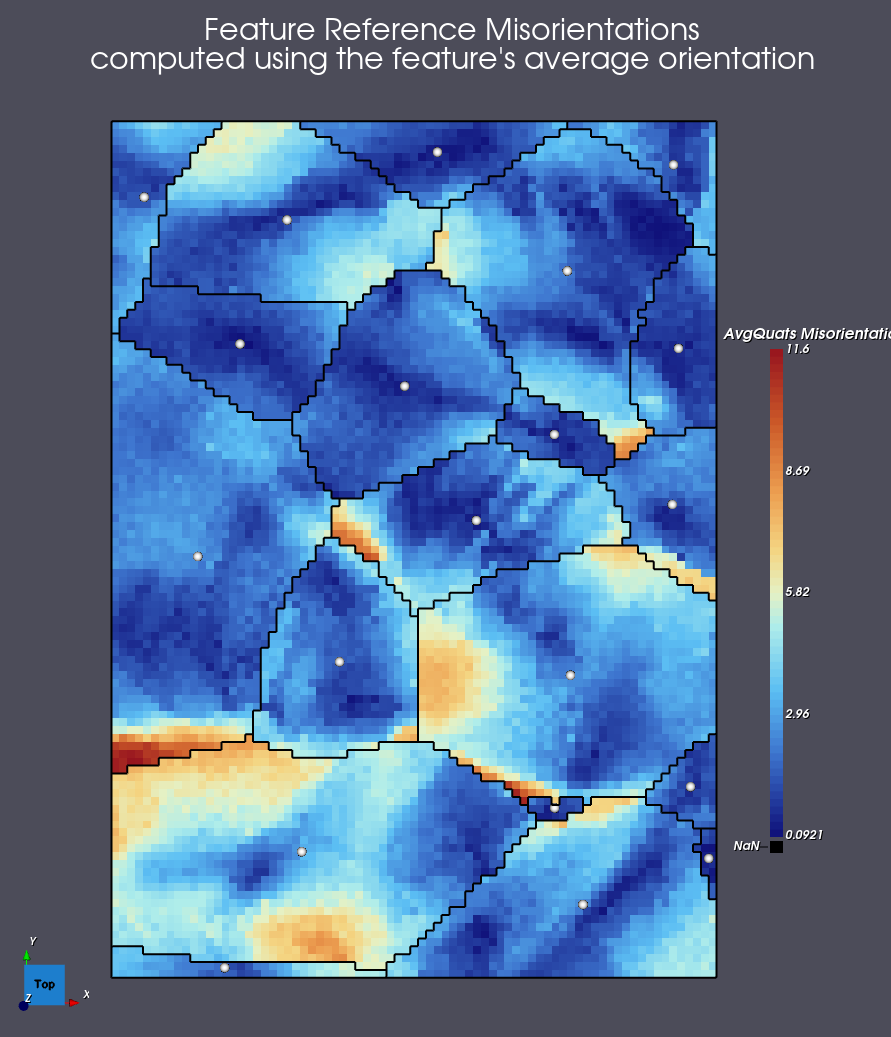
Example Using Feature’s Average Orientation
Using the T12-MAI-2010/fw-ar-IF1-aptr12-corr.ctf data set we can generate the following
data using the Feature Average Orientation choice.
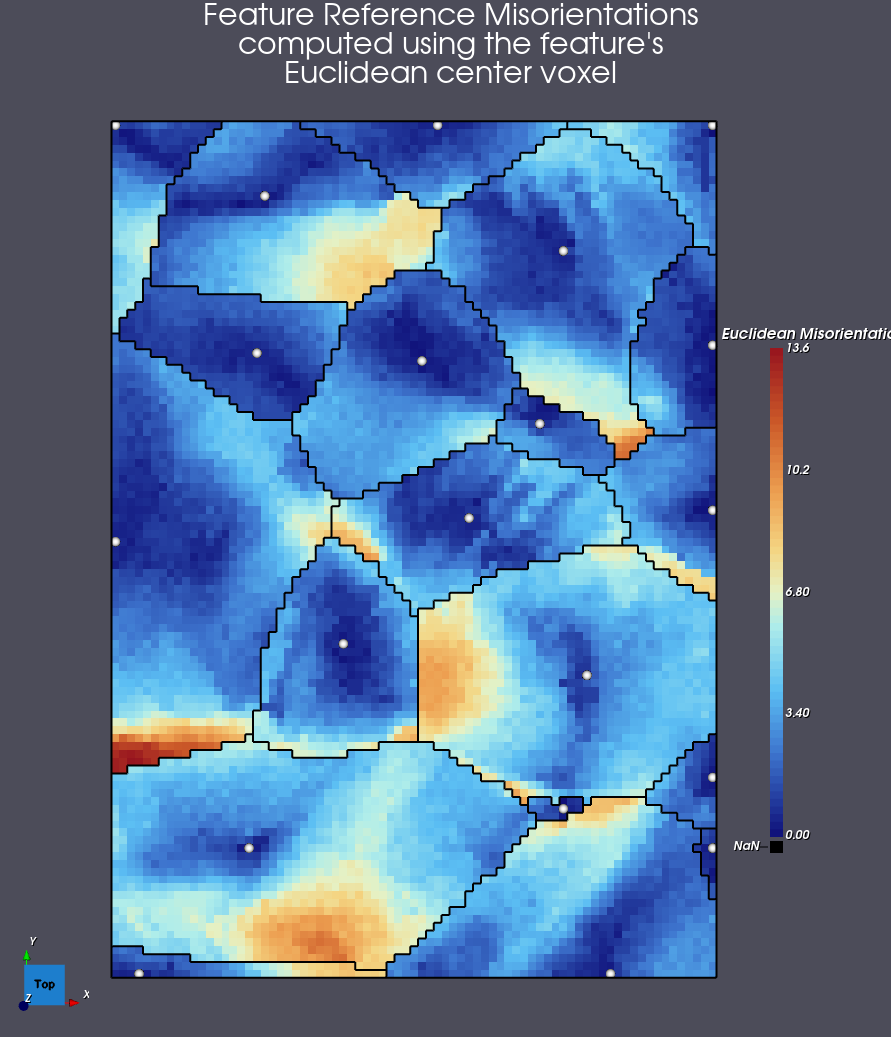
Example Using Feature’s Euclidean Center Voxel’s Orientation
Using the same dataset, the algorithm will find the voxel that is the furthest from the feature boundary, and use that voxel’s orientation as the reference orientation.
Input Parameter(s)
Parameter Name |
Parameter Type |
Parameter Notes |
Description |
|---|---|---|---|
Reference Orientation |
Choices |
Specifies the reference orientation to use when comparing to each Cell |
Input Cell Data
Parameter Name |
Parameter Type |
Parameter Notes |
Description |
|---|---|---|---|
Cell Feature Ids |
Array Selection |
Allowed Types: int32 Comp. Shape: 1 |
Specifies to which feature each cell belongs. |
Cell Phases |
Array Selection |
Allowed Types: int32 Comp. Shape: 1 |
Specifies to which Ensemble each Cell belongs |
Cell Quaternions |
Array Selection |
Allowed Types: float32 Comp. Shape: 4 |
Specifies the orientation of the Cell in quaternion representation |
Boundary Euclidean Distances |
Array Selection |
Allowed Types: float32 Comp. Shape: 1 |
Distance the Cells are from the boundary of the Feature they belong to. Only required if the reference orientation is selected to be the ‘Orientation Farthest from Feature Boundary’ |
Input Feature Data
Parameter Name |
Parameter Type |
Parameter Notes |
Description |
|---|---|---|---|
Average Quaternions |
Array Selection |
Allowed Types: float32 Comp. Shape: 4 |
Specifies the average orientation of the Feature in quaternion representation (, w). Only required if the reference orientation is selected to be the average of the Feature |
Feature Attribute Matrix |
AttributeMatrixSelection |
The path to the cell feature attribute matrix |
Input Ensemble Data
Parameter Name |
Parameter Type |
Parameter Notes |
Description |
|---|---|---|---|
Crystal Structures |
Array Selection |
Allowed Types: uint32 Comp. Shape: 1 |
Enumeration representing the crystal structure for each Ensemble |
Output Cell Data
Parameter Name |
Parameter Type |
Parameter Notes |
Description |
|---|---|---|---|
Cell Reference Misorientations |
DataObjectName |
The name of the array containing the misorientation angle (in degrees) between Cell’s orientation and the reference orientation of the Feature that owns that Cell |
Output Feature Data
Parameter Name |
Parameter Type |
Parameter Notes |
Description |
|---|---|---|---|
Feature Average Misorientations |
DataObjectName |
The name of the array containing the average of the Feature reference misorientation values for all of the Cells that belong to the Feature |
|
Feature Euclidean Centers |
DataObjectName |
The coordinate of the voxel that is used for the Euclidean based calculation |
Example Pipelines
(05) SmallIN100 Crystallographic Statistics
License & Copyright
Please see the description file distributed with this Plugin
DREAM3D-NX Help
If you need help, need to file a bug report or want to request a new feature, please head over to the DREAM3DNX-Issues GitHub site where the community of DREAM3D-NX users can help answer your questions.